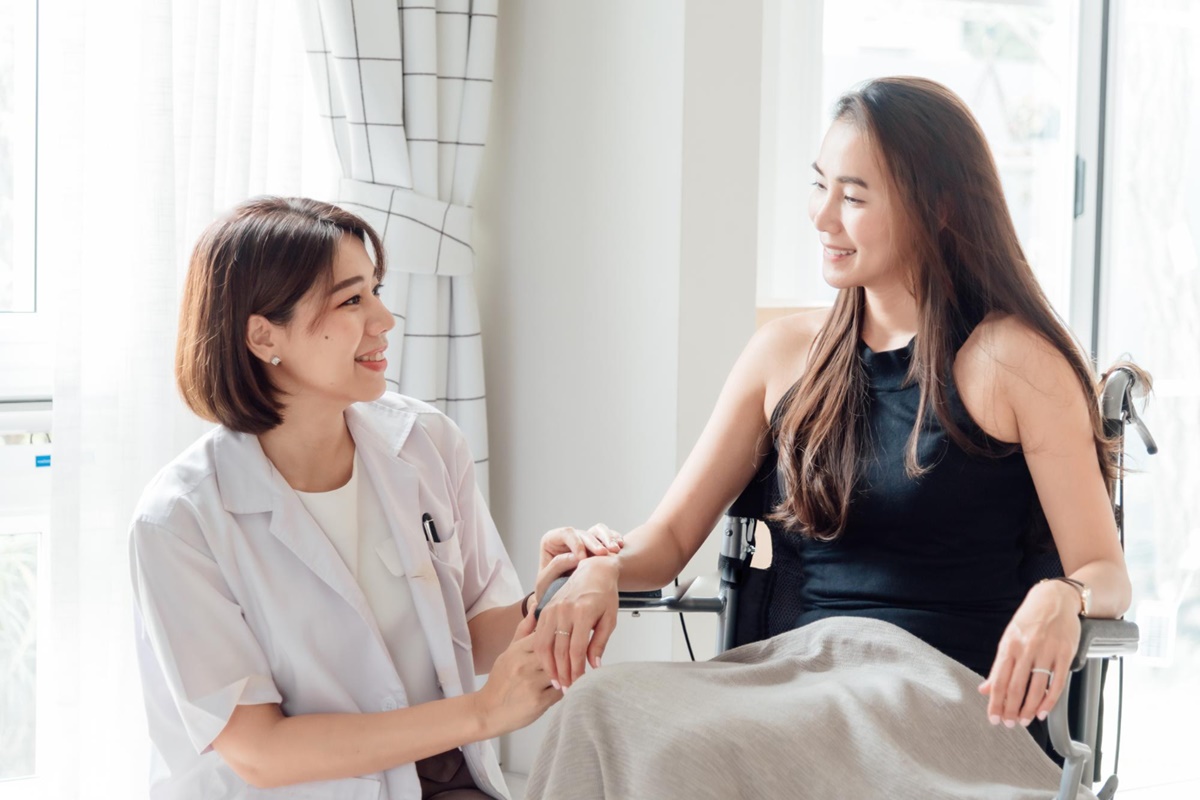
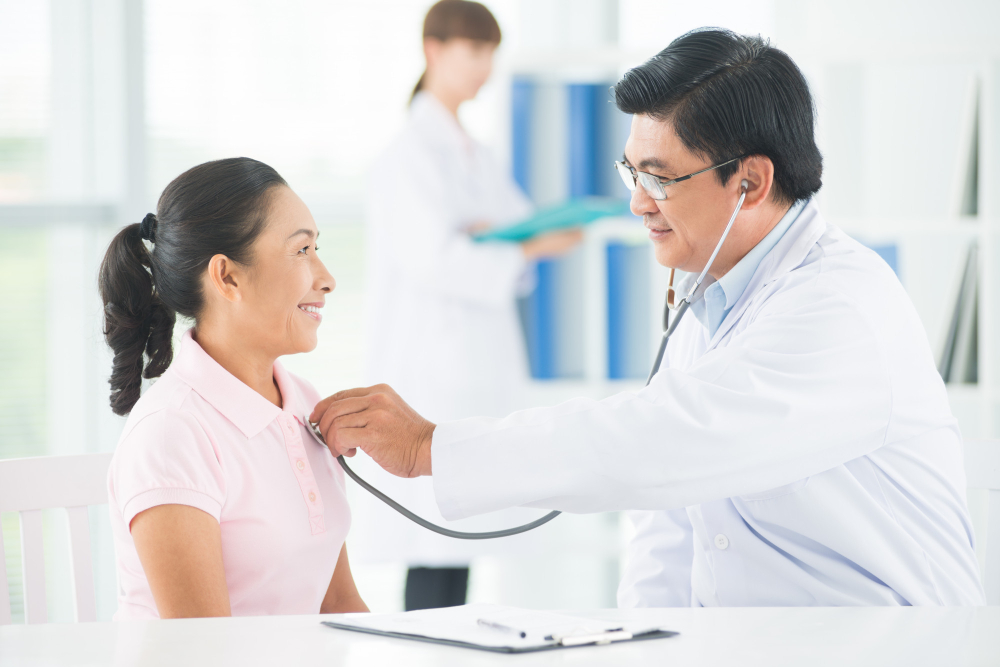
โรคออฟฟิศซินโดรม (Office Syndrome) เป็นโรคที่พบได้บ่อยในปัจจุบัน คลินิกกายภาพต่าง ๆ จึงมีโปรแกรมการรักษาโรคนี้ขึ้นมาโดยเฉพาะเพื่อช่วยบรรเทาอาการปวดของชาวออฟฟิศซินโดรมที่นั่งทำงานหรือนั่งท่าเดิมเป็นเวลานาน ๆ จนทำให้มีอาการปวดยึดตามกล้ามเนื้อส่วนต่าง ๆ เช่น บริเวณคอ บ่า หลัง แขน ข้อมือ ที่หากไม่รีบรักษาจะทำให้อาการปวดนั้นรุนแรงขึ้น ปวดเรื้อรัง ส่งผลต่อการทำงานและการใช้ชีวิตประจำวันได้ ...